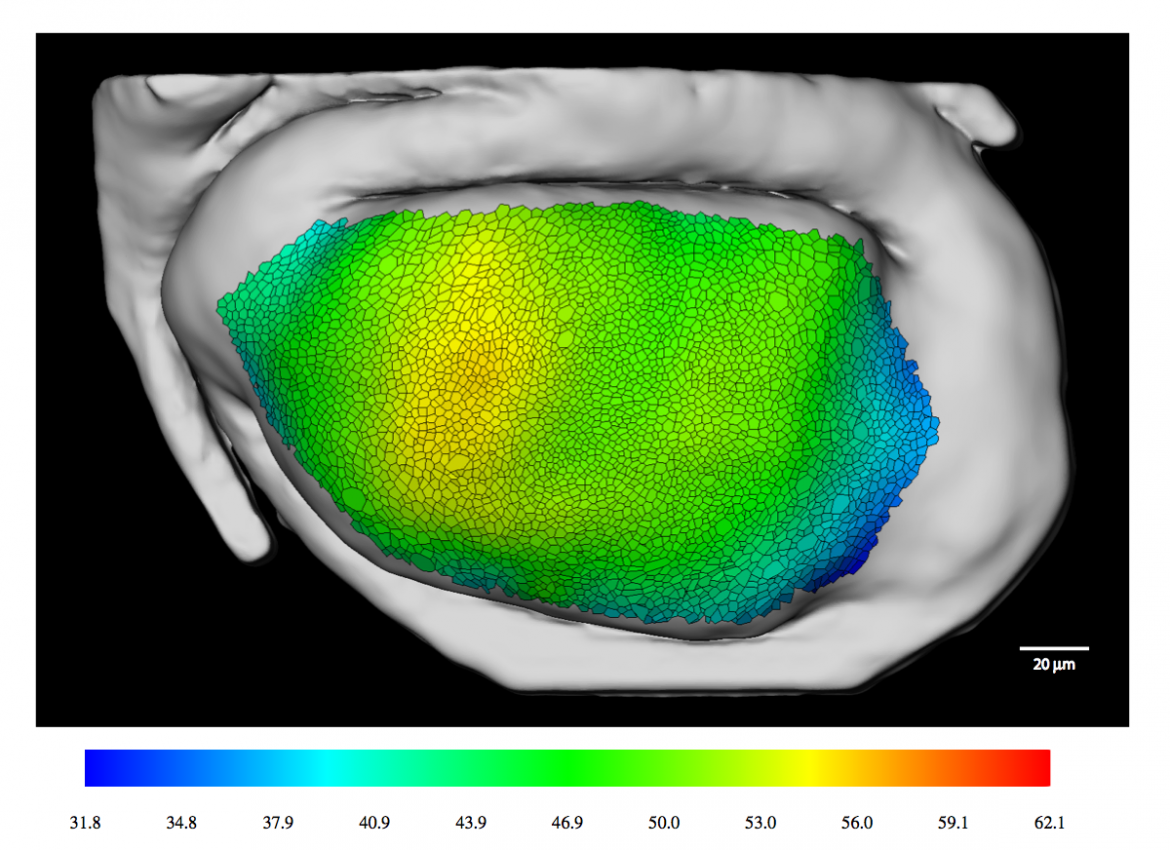
This is a novel and effective method for visualizing probabilistic tractograms within their anatomical context. This illustrative rendering technique, called fiber stippling, is inspired by visualization standards as found in anatomical textbooks. These illustrations typically show slice-based projections of fiber pathways and are typically hand-drawn. Applying the automatized technique to diffusion tractography, it is possible to demonstrate its expressiveness and intuitive usability as well as a more objective way to present white-matter structure in the human brain.
| Release Date: | |
| Status: | |
| Availability: | |
| Data type: | |
| Techniques: | |
| Software: | |
| Technology: | |
| Platform: | |
| Requirements: |
Project development
This work was supported by the German Federal Ministry of Education and Research as part of the VisPME research collaboration (01IH08009F) as well as by the AiF (ZIM grant KF 2034701SS8).
Last updated on 24th November, 2016
- Read more about OpenWalnut
- Log in or register to post comments